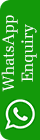Vol. 6, Issue 4, Part A (2019)
Immunoinformatics prediction for designing potential epitope-based malaria vaccine in the Aminopeptidase N1 protein of Anopheles gambiae (Diptera: Culicidae)
Author(s): Renu Jakhar¬, Pawan Kumar, Neelam Sehrawat and SK Gakhar
Abstract: Vector based Malaria Transmission Blocking Vaccines target antigens present on midgut surface of Anopheles mosquitoes. Aminopeptidase N is one of the important immunogenic protein involved in parasitic infection and its development in mosquitoes. In the present study, the epitopes that can encourage cell-mediated as well as humoral immunity against Anopheles gambiae Aminopeptidase N1 have been identified using immunoinformatics approach. Potential epitopes for cytotoxic T cells (MHCI) were identified using NetCTL and IEDB server. Selected T cell epitope were further analyzed by using computational docking technique to check interactions against the HLA molecules. Linear and conformational B cell epitope was also predicted and found to satisfy the threshold scores of all the parameters of predictive tools available on IEDB server. Therefore, an effort was done to design potential epitope based malaria vaccine by in silico analysis in the Aminopeptidase N1 protein of Anopheles gambiae.
 Fig.:
Fig.: HLA-C*14:02; Docking sites of predicted peptide (RRYLATTQF) against selected MHCI receptors.

How to cite this article:
Renu Jakhar¬, Pawan Kumar, Neelam Sehrawat, SK Gakhar. Immunoinformatics prediction for designing potential epitope-based malaria vaccine in the Aminopeptidase N1 protein of Anopheles gambiae (Diptera: Culicidae). Int J Mosq Res 2019;6(4):10-21.