Vol. 4, Issue 2, Part A (2017)
Molecular characterization of vitellogenesis in anautogenous Culex pipiens pipiens L. mosquitoes
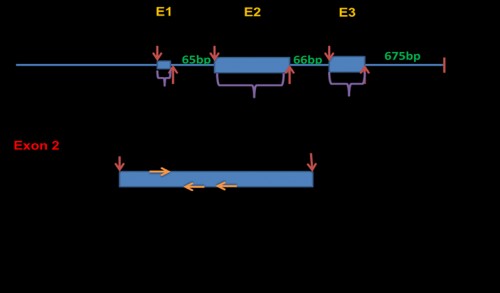
Fig. 1: Culex quinquefasciatus Vtg gene sequence diagram showing the intron-exon structure and the positions of the designed Vtg-specific primers (VgF, VgR1, VgR2) used for amplification of Vtg homologue(s) from in Cx. pipiens collected from Egypt. E: exon region.
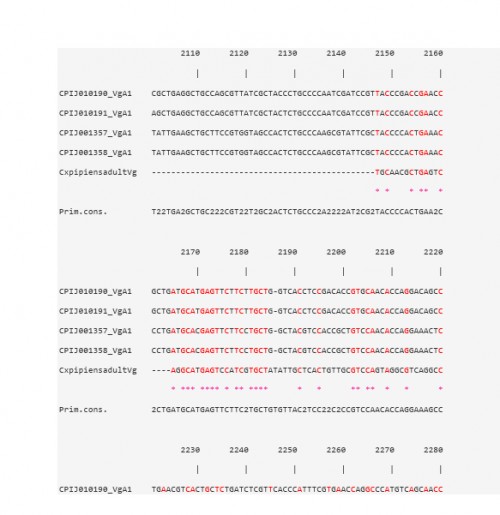
Fig. 2: CLUSTAL W multiple alignments at the Vtg nucleotide level of the sequence contig, for Cx. pipiens females 48 h PBM, with four Vg-A1 transcripts in Cx. quinquefasciatus

Fig. 3: CLUSTAL W multiple alignments at the Vtg nucleotide level of the sequence contig, for Cx. pipiens females 48 h PBM, with four Vg-A1 transcripts in Cx. quinquefasciatus
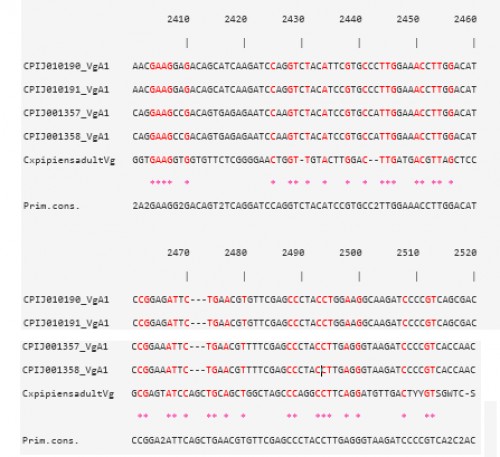
Fig. 4: CLUSTAL W multiple alignments at the Vtg nucleotide level of the sequence contig, for Cx. pipiens females 48 h PBM, with four Vg-A1 transcripts in Cx. quinquefasciatus

Fig. 5: CLUSTAL W multiple alignments at the Vtg nucleotide level of the sequence contig, for Cx. pipiens females 48 h PBM, with four Vg-A1 transcripts in Cx. quinquefasciatus
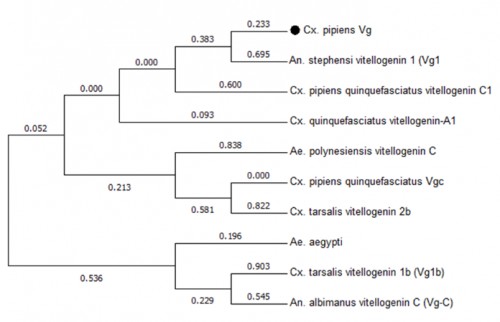
Fig. 6: Phylogenetic analysis of Vtg gene transcripts from various species of mosquitoes. The analysis involved 10 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 39 positions in the final dataset
Fig. 7: Quantitative expression of Cx. pipiens Vtg gene in immatures and adult mosquitoes at different feeding status. Each point represents the average of three replicates. Bars denote to the standard error
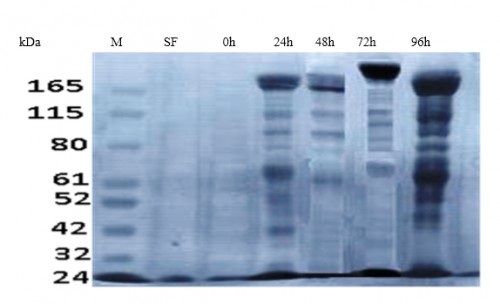
Fig. 8: SDS-PAGE of total Coomassie Brilliant Blue stained ovarian protein extract (20 mg/ ml). SF: sugar-fed females, 0: immediately PBM, 24 h PBM, 48 h PBM, 72 h PBM, 96 h PBM, M: protein ladder (KDa)